Performing Tritium Breeding Ratio (TBR) simulations#
This python notebook allows users to perform neutronics simulations to tally tritium production.
This example uses a simple sphere model with a breeder material and a tritium production tally.
First import OpenMC and configure the nuclear data path
import openmc
from pathlib import Path
# Setting the cross section path to the correct location in the docker image.
# If you are running this outside the docker image you will have to change this path to your local cross section path.
openmc.config['cross_sections'] = Path.home() / 'nuclear_data' / 'cross_sections.xml'
This first code block makes the geometry, materials and settings for the neutronics model.
# MATERIALS
breeder_material = openmc.Material() # lithium lead chemical equation is Pb84.2Li15.8
breeder_material.add_element('Pb', 84.2, percent_type='ao')
# 50% enriched lithium 6, natural percentage of lithium 6 is just 7%
breeder_material.add_element('Li', 15.8, percent_type='ao', enrichment=50.0, enrichment_target='Li6', enrichment_type='ao')
# we set density using atoms per b-cm units as then when the enrichment changes the dnesity in these units stays the same.
# It equates to around 11 g/cm3
breeder_material.set_density('atom/b-cm', 3.2720171e-2)
steel = openmc.Material()
steel.set_density('g/cm3', 7.75)
steel.add_element('Fe', 0.95, percent_type='wo')
steel.add_element('C', 0.05, percent_type='wo')
my_materials = openmc.Materials([breeder_material, steel])
# GEOMETRY
# surfaces
vessel_inner = openmc.Sphere(r=500)
first_wall_outer_surface = openmc.Sphere(r=510)
breeder_blanket_outer_surface = openmc.Sphere(r=610, boundary_type='vacuum')
# cells
inner_vessel_region = -vessel_inner
inner_vessel_cell = openmc.Cell(region=inner_vessel_region)
first_wall_region = -first_wall_outer_surface & +vessel_inner
first_wall_cell = openmc.Cell(region=first_wall_region)
first_wall_cell.fill = steel
breeder_blanket_region = +first_wall_outer_surface & -breeder_blanket_outer_surface
breeder_blanket_cell = openmc.Cell(region=breeder_blanket_region)
breeder_blanket_cell.fill = breeder_material
my_geometry = openmc.Geometry([inner_vessel_cell, first_wall_cell, breeder_blanket_cell])
# SIMULATION SETTINGS
# Instantiate a Settings object
my_settings = openmc.Settings()
my_settings.batches = 10
my_settings.particles = 500
my_settings.run_mode = 'fixed source'
# Create a DT point source
my_source = openmc.IndependentSource(
space=openmc.stats.Point((0, 0, 0)),
angle=openmc.stats.Isotropic(),
energy=openmc.stats.Discrete([14e6], [1])
)
my_settings.source = my_source
# added a cell tally for tritium production
cell_filter = openmc.CellFilter(breeder_blanket_cell)
tbr_tally = openmc.Tally(name='TBR')
tbr_tally.filters = [cell_filter]
tbr_tally.scores = ['H3-production'] # Equivalent to '(n,Xt)': X is a wildcard for outgoing neutron count, t = triton (tritium nucleus). Covers both (n,t) and (n,nt) reactions in Li6/Li7.
# this allows the tally to be recorded per nuclide so we can see which one contributes to tritium production more
tbr_tally.nuclides = ['Li6', 'Li7']
my_tallies = openmc.Tallies([tbr_tally])
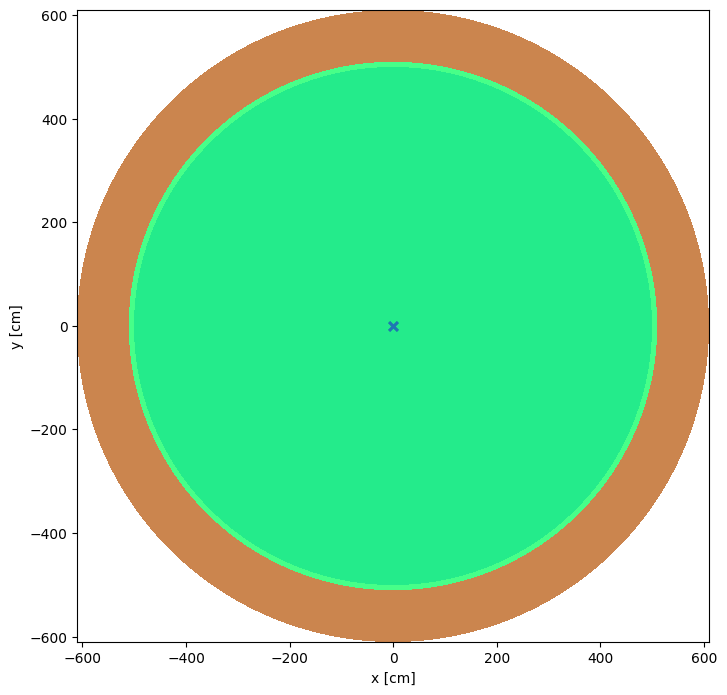
Now we assembly the model and plot it just to check we have made the model the way we thing that we have.
model = openmc.Model(my_geometry, my_materials, my_settings, my_tallies)
model.plot(n_samples=10, pixels=400000)
<Axes: xlabel='x [cm]', ylabel='y [cm]'>
The next code block runs the simulation.
# removes the old output files
for f in Path('.').glob('*.h5'):
f.unlink(missing_ok=True)
# Run OpenMC!
sp_filename = model.run()
%%%%%%%%%%%%%%%
%%%%%%%%%%%%%%%%%%%%%%%%
%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
%%%%%%%%%%%%%%%%%%%%%%%%
%%%%%%%%%%%%%%%%%%%%%%%%
############### %%%%%%%%%%%%%%%%%%%%%%%%
################## %%%%%%%%%%%%%%%%%%%%%%%
################### %%%%%%%%%%%%%%%%%%%%%%%
#################### %%%%%%%%%%%%%%%%%%%%%%
##################### %%%%%%%%%%%%%%%%%%%%%
###################### %%%%%%%%%%%%%%%%%%%%
####################### %%%%%%%%%%%%%%%%%%
####################### %%%%%%%%%%%%%%%%%
###################### %%%%%%%%%%%%%%%%%
#################### %%%%%%%%%%%%%%%%%
################# %%%%%%%%%%%%%%%%%
############### %%%%%%%%%%%%%%%%
############ %%%%%%%%%%%%%%%
######## %%%%%%%%%%%%%%
%%%%%%%%%%%
| The OpenMC Monte Carlo Code
Copyright | 2011-2026 MIT, UChicago Argonne LLC, and contributors
License | https://docs.openmc.org/en/latest/license.html
Version | 0.15.3-dev485
Commit Hash | 6704e4786a743611baac5d693943f67f7479ee47
Date/Time | 2026-04-02 12:57:28
OpenMP Threads | 4
Reading model XML file 'model.xml' ...
Reading cross sections XML file...
Reading Pb204 from /home/runner/nuclear_data/neutron/Pb204.h5
Reading Pb206 from /home/runner/nuclear_data/neutron/Pb206.h5
Reading Pb207 from /home/runner/nuclear_data/neutron/Pb207.h5
Reading Pb208 from /home/runner/nuclear_data/neutron/Pb208.h5
Reading Li6 from /home/runner/nuclear_data/neutron/Li6.h5
Reading Li7 from /home/runner/nuclear_data/neutron/Li7.h5
Reading Fe54 from /home/runner/nuclear_data/neutron/Fe54.h5
Reading Fe56 from /home/runner/nuclear_data/neutron/Fe56.h5
Reading Fe57 from /home/runner/nuclear_data/neutron/Fe57.h5
Reading Fe58 from /home/runner/nuclear_data/neutron/Fe58.h5
Reading C12 from /home/runner/nuclear_data/neutron/C12.h5
Reading C13 from /home/runner/nuclear_data/neutron/C13.h5
Minimum neutron data temperature: 294.0 K
Maximum neutron data temperature: 294.0 K
Preparing distributed cell instances...
Writing summary.h5 file...
Maximum neutron transport energy: 20000000.0 eV for Li6
===============> FIXED SOURCE TRANSPORT SIMULATION <===============
Simulating batch 1
Simulating batch 2
Simulating batch 3
Simulating batch 4
Simulating batch 5
Simulating batch 6
Simulating batch 7
Simulating batch 8
Simulating batch 9
Simulating batch 10
Creating state point statepoint.10.h5...
=======================> TIMING STATISTICS <=======================
Total time for initialization = 2.0713e+00 seconds
Reading cross sections = 2.0603e+00 seconds
Total time in simulation = 3.3937e-01 seconds
Time in transport only = 3.3367e-01 seconds
Time in active batches = 3.3937e-01 seconds
Time accumulating tallies = 3.9544e-05 seconds
Time writing statepoints = 4.6240e-03 seconds
Total time for finalization = 1.3449e-04 seconds
Total time elapsed = 2.4140e+00 seconds
Calculation Rate (active) = 14733.3 particles/second
============================> RESULTS <============================
Leakage Fraction = 0.04373 +/- 0.00282
Finally, this code block loads up the output file and extracts the simulation result.
# open the results file
sp = openmc.StatePoint(sp_filename)
# access the tally using pandas dataframes
tbr_tally = sp.get_tally(name='TBR')
df = tbr_tally.get_pandas_dataframe()
# prints the contents of the dataframe
df
| cell | nuclide | score | mean | std. dev. | |
|---|---|---|---|---|---|
| 0 | 3 | Li6 | (n,Xt) | 1.022777 | 0.018036 |
| 1 | 3 | Li7 | (n,Xt) | 0.003777 | 0.000164 |
We can access the values from the dataframe
# sums up all the values in the mean column
tbr_tally_result = df['mean'].sum()
# sums up all the values in the std. dev. column
tbr_tally_std_dev = df['std. dev.'].sum()
# print the results
print('The tritium breeding ratio was found, TBR = ', tbr_tally_result)
print('Standard deviation on the tbr tally is ', tbr_tally_std_dev)
The tritium breeding ratio was found, TBR = 1.026554366872371
Standard deviation on the tbr tally is 0.0181998968207567
We can also access the individual results of tritium production in each isotope. This is provided because we added ```.nucl
Learning Outcomes for Part 1:
Running simple neutronics simulations with OpenMC and tallying TBR using a cell tally.
# gets the row that has li6 in the nuclide column and
lithium_6_contribution = df[df['nuclide'] == 'Li6']['mean'].sum()
lithium_7_contribution = df[df['nuclide'] == 'Li7']['mean'].sum()
print('The tritium breeding by lithium 6 is ', lithium_6_contribution)
print('The tritium breeding by lithium 7 is ', lithium_7_contribution)
The tritium breeding by lithium 6 is 1.022777369594151
The tritium breeding by lithium 7 is 0.0037769972782197908
Learning Outcomes for Part 1:
Running simple neutronics simulations with OpenMC and tallying TBR using a cell tally.
Identify the which lithium isotope makes the majority of tritium production

